|
ADLEMAN L M. Molecular computation of solutions to combinatorial problems[J]. Science, 1994, 266(5187): 1021–1024. doi: 10.1126/science.7973651
|
|
王雷, 林亚平. DNA计算在整数规划问题中的应用[J]. 电子与信息学报, 2005, 27(5): 814–818.WANG Lei and LIN Yaping. DNA computation for a category of special integer planning problem[J]. Journal of Electronics &Information Technology, 2005, 27(5): 814–818.
|
|
吴雪, 赵艺. 最大加权独立集问题的DNA算法[J]. 电子与信息学报, 2007, 29(11): 2693–2697.WU Xue and ZHAO Yi. DNA solution of the maximum weighted independent set[J]. Journal of Electronics &Information Technology, 2007, 29(11): 2693–2697.
|
|
KALLENBACH N R, MA R I, and SEEMAN N C. An immobile nucleic acid junction constructed from oligonucleotides[J]. Nature, 1983, 305(5937): 829–831. doi: 10.1038/305829a0
|
|
FU T J and SEEMAN N C. DNA double-crossover molecules[J]. Biochemistry, 1992, 32(13): 3211–3220. doi: 10.1021/bi00064a003
|
|
WINFREE E, LIU Furong, WENZLER L A, et al. Design and self-assembly of two-dimensional DNA crystals[J]. Nature, 1998, 394(6693): 539–544. doi: 10.1038/28998
|
|
YAN Hao, PARK S H, FINKELSTEIN G, et al. DNA-templated self-assembly of protein arrays and highly conductive nanowires[J]. Science, 2003, 301(5641): 1882–1884. doi: 10.1126/science.1089389
|
|
WEI B, DAI Mingjie, and YIN Peng. Complex shapes self-assembled from single-stranded DNA tiles[J]. Nature, 2012, 485(7400): 623–626. doi: 10.1038/nature11075
|
|
SHI Xiaolong, LU Wei, WANG Zhiyu, et al. Programmable DNA tile self-assembly using a hierarchical sub-tile strategy[J]. Nanotechnology, 2014, 25(7): 075602. doi: 10.1088/0957-4484/25/7/075602
|
|
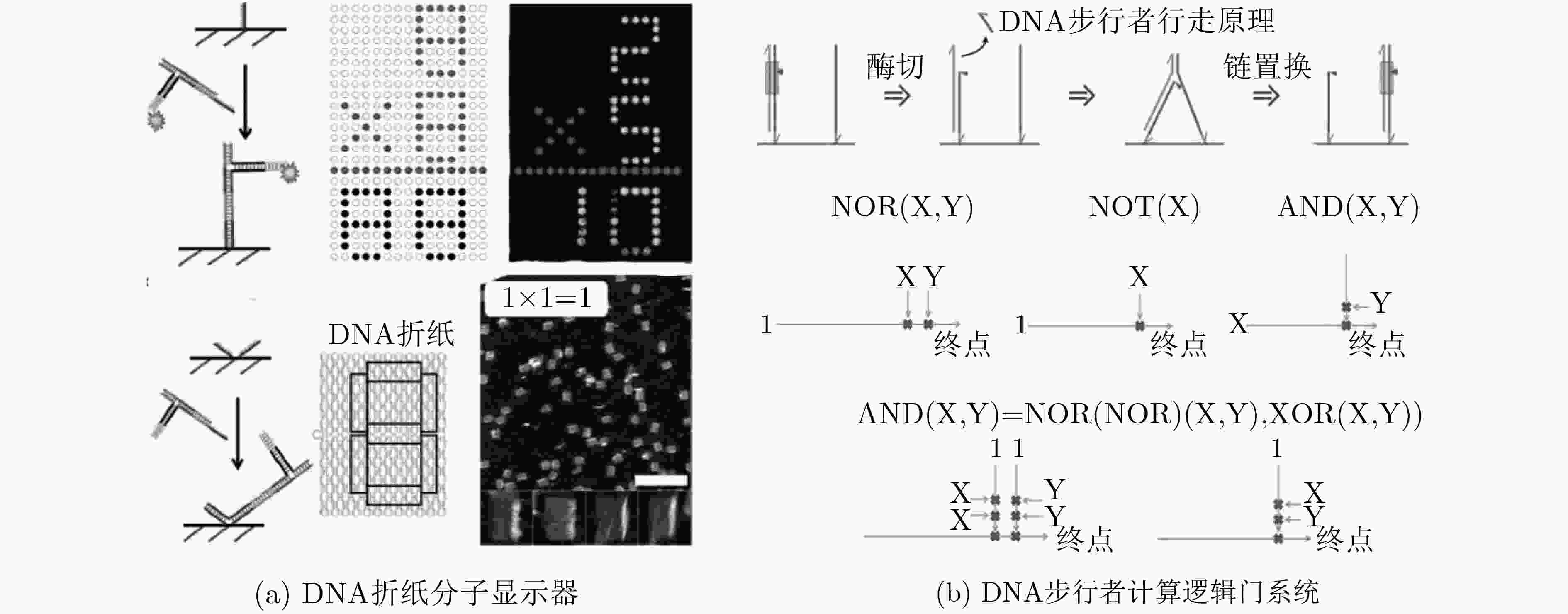
ROTHEMUND P W K. Folding DNA to create nanoscale shapes and patterns[J]. Nature, 2006, 440(7082): 297–302. doi: 10.1038/nature04586
|
|
QIAN Lulu, WANG Ying, ZHANG Zhao, et al. Analogic China map constructed by DNA[J]. Chinese Science Bulletin, 2006, 51(24): 2973–2976. doi: 10.1007/s11434-006-2223-9
|
|
ANDERSEN E S, DONG Mingdong, NIELSEN M M, et al. DNA origami design of dolphin-shaped structures with flexible tails[J]. ACS Nano, 2008, 2(6): 1213–1218. doi: 10.1021/nn800215j
|
|
ZHANG Honglu, CHAO Jie, PAN Dun, et al. Folding super-sized DNA origami with scaffold strands from long-range PCR[J]. Chemical Communications, 2012, 48(51): 6405–6407. doi: 10.1039/c2cc32204h
|
|
DOUGLAS S M, DIETZ H, LIEDL T, et al. Self-assembly of DNA into nanoscale three-dimensional shapes[J]. Nature, 2009, 459(7245): 414–418. doi: 10.1038/nature08016
|
|
DOUGLAS S M, MARBLESTONE A H, TEERAPITTAYANON S, et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno[J]. Nucleic Acids Research, 2009, 37(15): 5001–5006. doi: 10.1093/nar/gkp436
|
|
DIETZ H, DOUGLAS S M, and SHIH W M. Folding DNA into twisted and curved nanoscale shapes[J]. Science, 2009, 325(5941): 725–730. doi: 10.1126/science.1174251
|
|
KIM Y J, LEE C, LEE J G, et al. Configurational design of mechanical perturbation for fine control of twisted dna origami structures[J]. ACS Nano, 2019, 13(6): 6348–6355. doi: 10.1021/acsnano.9b01561
|
|
HAN Dongran, PAL S, NANGREAVE J, et al. DNA origami with complex curvatures in three-dimensional space[J]. Science, 2011, 332(6027): 342–346. doi: 10.1126/science.1202998
|
|
LIN Zhiwei, XIONG Yan, XIANG Shuting, et al. Controllable covalent-bound nanoarchitectures from DNA frames[J]. Journal of the American Chemical Society, 2019, 141(17): 6797–6801. doi: 10.1021/jacs.9b01510
|
|
ZHANG Tao, HARTL C, FRANK K, et al. 3D DNA origami crystals[J]. Advanced Materials, 2018, 30(28): 1800273. doi: 10.1002/adma.201800273
|
|
HAN Dongran, PAL S, YANG Yang, et al. DNA gridiron nanostructures based on four-arm junctions[J]. Science, 2013, 339(6126): 1412–1415. doi: 10.1126/science.1232252
|
|
KE Yonggang, ONG L L, SUN Wei, et al. DNA brick crystals with prescribed depths[J]. Nature Chemistry, 2014, 6(11): 994–1002. doi: 10.1038/nchem.2083
|
|
WANG Wen, CHEN Silian, AN B, et al. Complex wireframe DNA nanostructures from simple building blocks[J]. Nature Communications, 2019, 10(1): 1067. doi: 10.1038/s41467-019-08647-7
|
|
STOJANOVIC M N, MITCHELL T E, and STEFANOVIC D. Deoxyribozyme-based logic gates[J]. Journal of the American Chemical Society, 2002, 124(14): 3555–3561. doi: 10.1021/ja016756v
|
|
STOJANOVIC M N and STEFANOVIC D. Deoxyribozyme-based half-adder[J]. Journal of the American Chemical Society, 2003, 125(22): 6673–6676. doi: 10.1021/ja0296632
|
|
PENCHOVSKY R and BREAKER R R. Computational design and experimental validation of oligonucleotide-sensing allosteric ribozymes[J]. Nature Biotechnology, 2005, 23(11): 1424–1433. doi: 10.1038/nbt1155
|
|
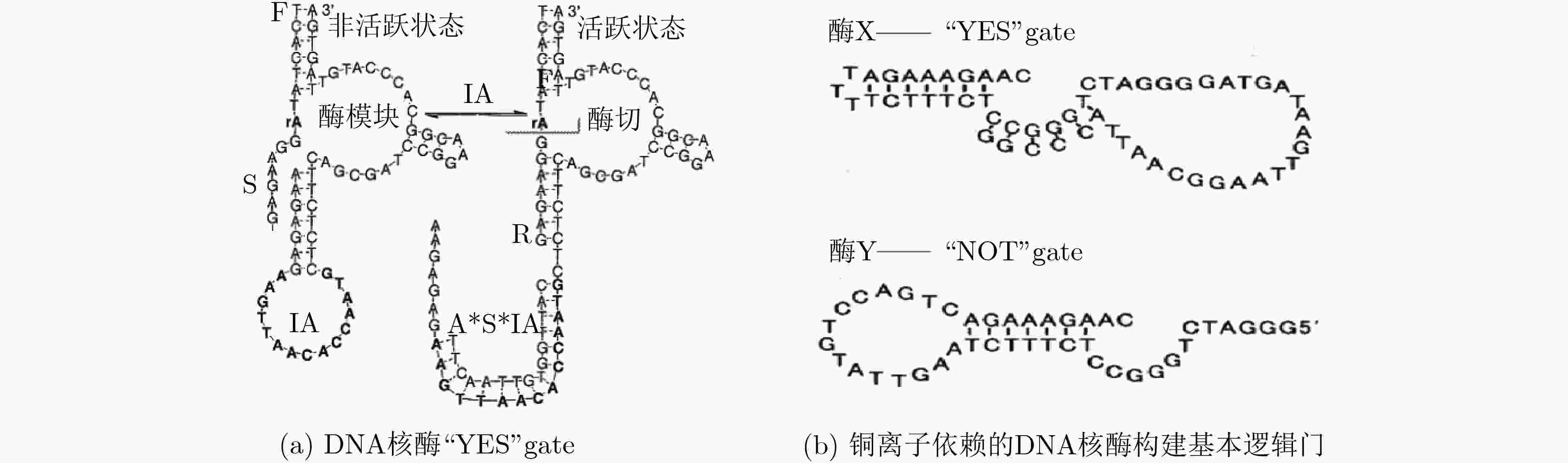
CHEN Xi, WANG Yifei, LIU Qiang, et al. Construction of molecular logic gates with a DNA-cleaving deoxyribozyme[J]. Angewandte Chemie, 2006, 118(11): 1791–1794. doi: 10.1002/ange.200502511
|
|
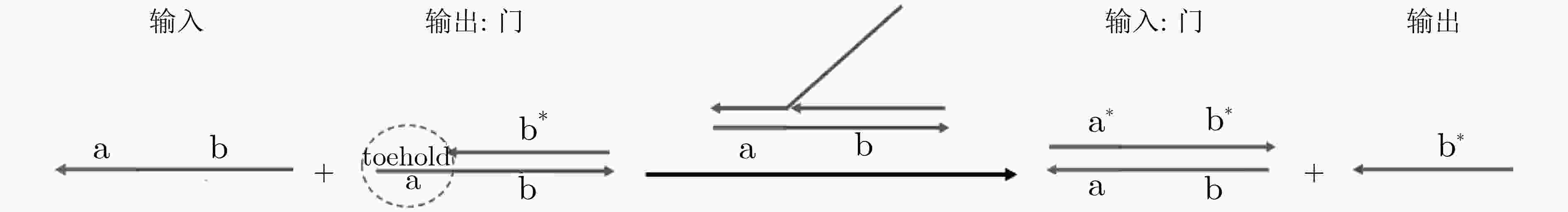
SEELIG G, SOLOVEICHIK D, ZHANG D Y, et al. Enzyme-free nucleic acid logic circuits[J]. Science, 2006, 314(5805): 1585–1588. doi: 10.1126/science.1132493
|
|
QIAN Lulu and WINFREE E. A simple DNA gate motif for synthesizing large-scale circuits[J]. Journal of the Royal Society Interface, 2011, 8(62): 1281–1297. doi: 10.1098/rsif.2010.0729
|
|
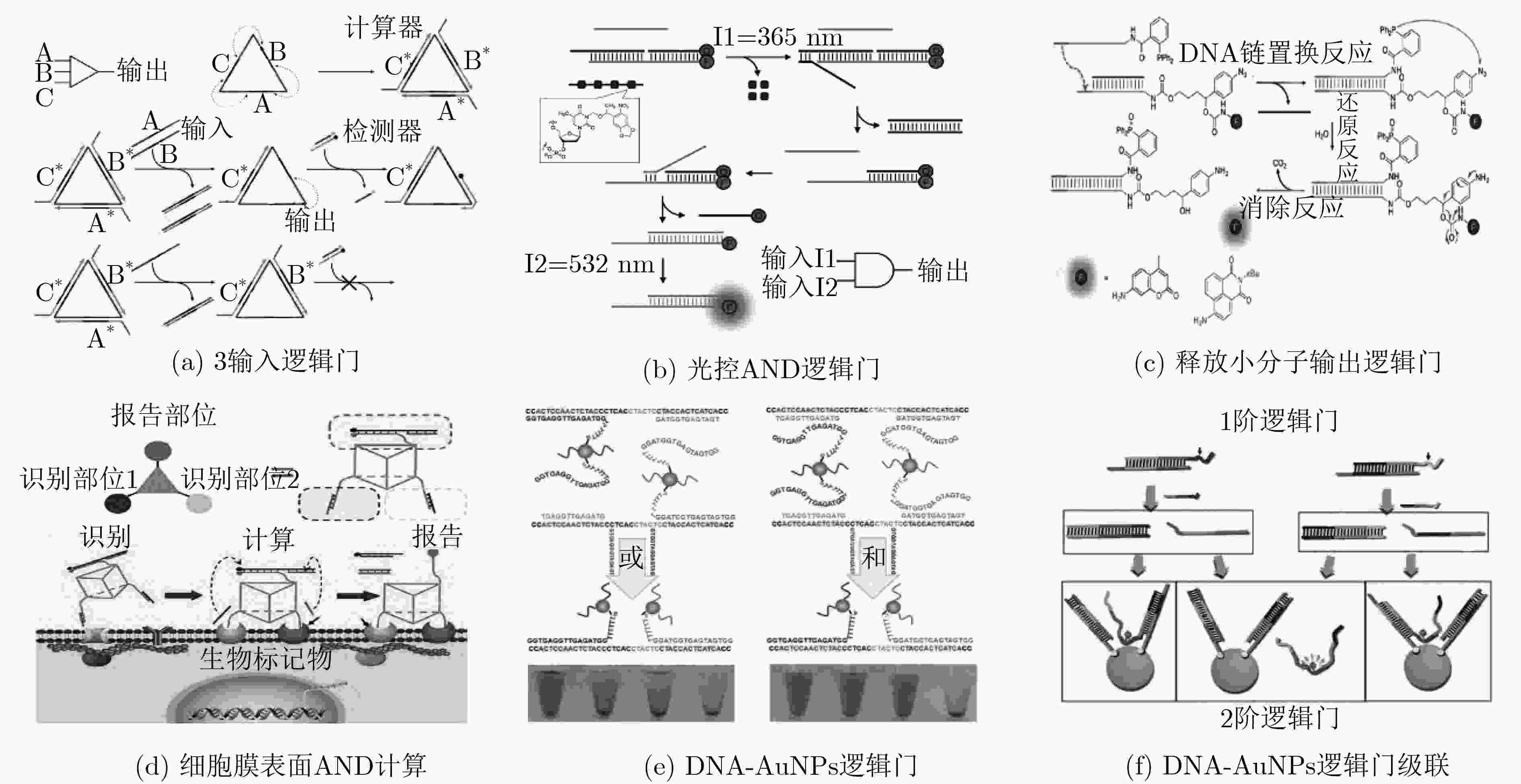
LI Wei, YANG Yang, YAN Hao, et al. Three-input majority logic gate and multiple input logic circuit based on DNA strand displacement[J]. Nano Letters, 2013, 13(6): 2980–2988. doi: 10.1021/nl4016107
|
|
PROKUP A, HEMPHILL J, and DEITERS A. DNA computation: A photochemically controlled and gate[J]. Journal of the American Chemical Society, 2012, 134(8): 3810–3815. doi: 10.1021/ja210050s
|
|
HEMPHILL J and DEITERS A. DNA computation in mammalian cells: MicroRNA logic operations[J]. Journal of the American Chemical Society, 2013, 135(28): 10512–10518. doi: 10.1021/ja404350s
|
|
MORIHIRO K, ANKENBRUCK N, LUKASAK B, et al. Small molecule release and activation through DNA computing[J]. Journal of the American Chemical Society, 2017, 139(39): 13909–13915. doi: 10.1021/jacs.7b07831
|
|
PENG Ruizi, ZHENG Xiaofang, LYU Yifan, et al. Engineering a 3D DNA-logic gate nanomachine for bispecific recognition and computing on target cell surfaces[J]. Journal of the American Chemical Society, 2018, 140(31): 9793–9796. doi: 10.1021/jacs.8b04319
|
|
SONG Tingjie and LIANG Haojun. Synchronized assembly of gold nanoparticles driven by a dynamic DNA-fueled molecular machine[J]. Journal of the American Chemical Society, 2012, 134(26): 10803–10806. doi: 10.1021/ja304746k
|
|
YANG Jing, SHEN Lingjing, MA Jingjing, et al. Fluorescent nanoparticle beacon for logic gate operation regulated by strand displacement[J]. ACS Applied Materials & Interfaces, 2013, 5(12): 5392–5396. doi: 10.1021/am401493d
|
|
NIKITIN M P, SHIPUNOVA V O, DEYEV S M, et al. Biocomputing based on particle disassembly[J]. Nature Nanotechnology, 2014, 9(9): 716–722. doi: 10.1038/nnano.2014.156
|
|
PHILLIPS A and CARDELLI L. A programming language for composable DNA circuits[J]. Journal of the Royal Society Interface, 2009, 6(Suppl 4): S419–S436. doi: 10.1098/rsif.2009.0072.focus
|
|
LAKIN M R, YOUSSEF S, POLO F, et al. Visual DSD: A design and analysis tool for DNA strand displacement systems[J]. Bioinformatics, 2011, 27(22): 3211–3213. doi: 10.1093/bioinformatics/btr543
|
|
GEORGE A K and SINGH H. DNA strand displacement-based logic inverter gate design[J]. Micro & Nano Letters, 2017, 12(9): 611–614. doi: 10.1049/mnl.2017.0142
|
|
NIU YING, SHEN CHAONAN, and ZHANG XUNCAI. Design of logic circuits based on metallo-toehold strand displacement[J]. Journal of Nanoelectronics and Optoelectronics, 2019, 14(2): 232–237. doi: 10.1166/jno.2019.2480
|
|
CHERRY K M and QIAN Lulu. Scaling up molecular pattern recognition with DNA-based winner-take-all neural networks[J]. Nature, 2018, 559(7714): 370–376. doi: 10.1038/s41586-018-0289-6
|
|
ANDERSEN E S, DONG Mingdong, NIELSEN M M, et al. Self-assembly of a nanoscale DNA box with a controllable lid[J]. Nature, 2009, 459(7243): 73–76. doi: 10.1038/nature07971
|
|
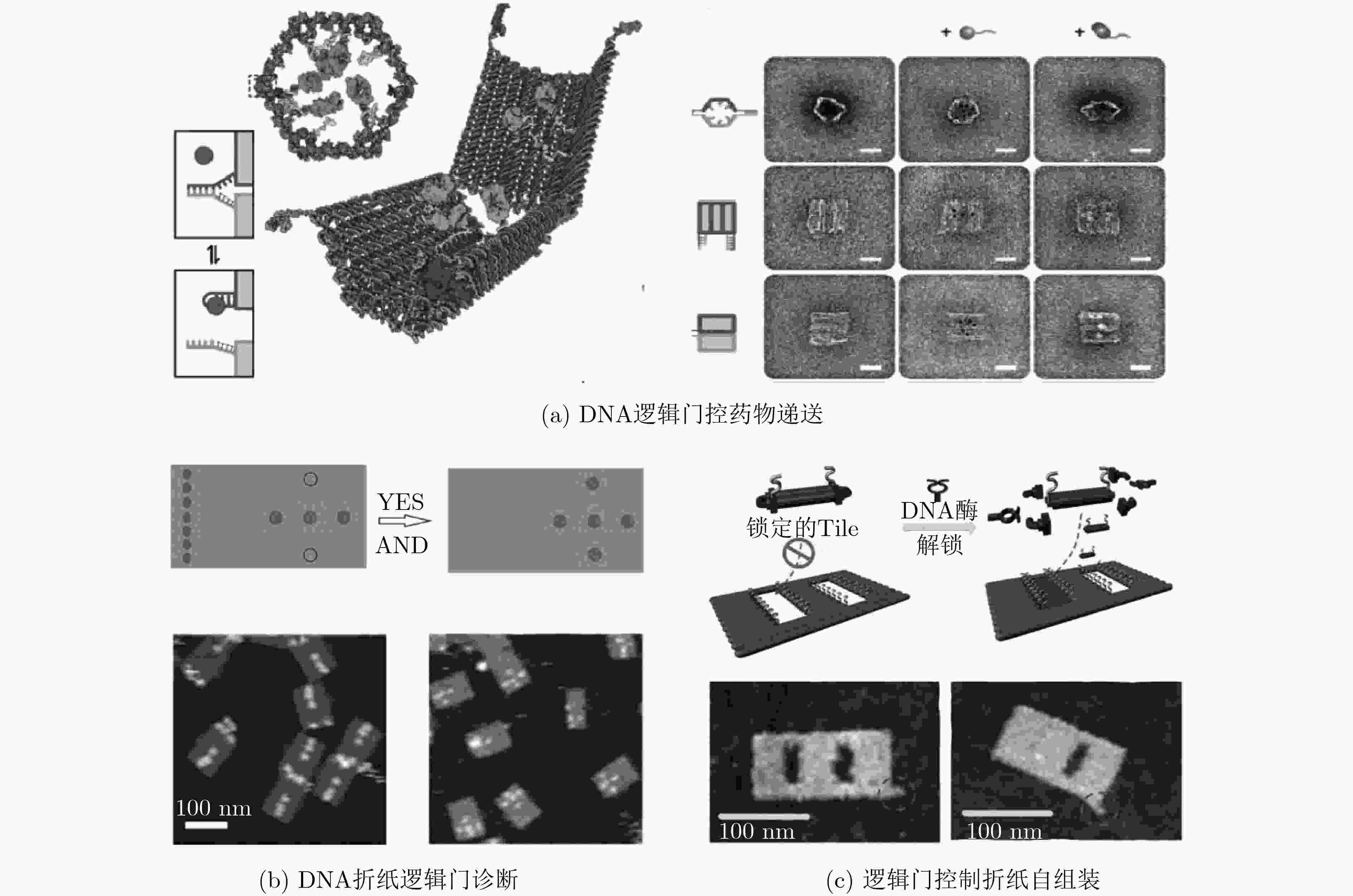
DOUGLAS S M, BACHELET I, and CHURCH G M. A logic-gated nanorobot for targeted transport of molecular payloads[J]. Science, 2012, 335(6070): 831–834. doi: 10.1126/science.1214081
|
|
AMIR Y, BEN-ISHAY E, LEVNER D, et al. Universal computing by DNA origami robots in a living animal[J]. Nature Nanotechnology, 2014, 9(5): 353–357. doi: 10.1038/nnano.2014.58
|
|
WANG Dongfang, FU Yanming, YAN Juan, et al. Molecular logic gates on DNA origami nanostructures for microRNA diagnostics[J]. Analytical Chemistry, 2014, 86(4): 1932–1936. doi: 10.1021/ac403661z
|
|
ZHANG Cheng, YANG Jing, JIANG Shuoxing, et al. DNAzyme-based logic gate-mediated DNA self-assembly[J]. Nano Letters, 2016, 16(1): 736–741. doi: 10.1021/acs.nanolett.5b04608
|
|
STOJANOVIC M N and STEFANOVIC D. A deoxyribozyme-based molecular automaton[J]. Nature Biotechnology, 2003, 21(9): 1069–1074. doi: 10.1038/nbt862
|
|
MACDONALD J, LI Yang, SUTOVIC M, et al. Medium scale integration of molecular logic gates in an automaton[J]. Nano Letters, 2006, 6(11): 2598–2603. doi: 10.1021/nl0620684
|
|
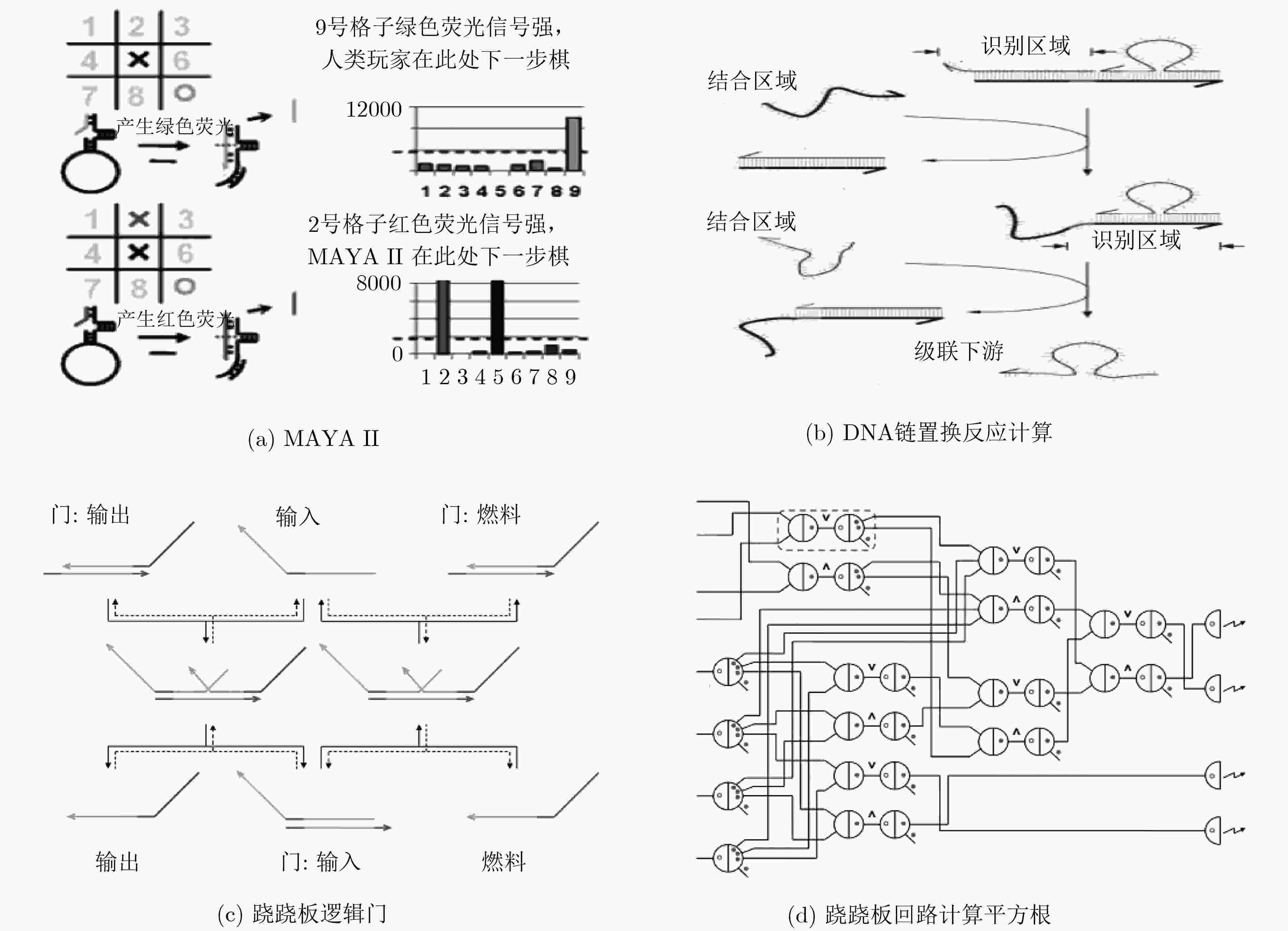
PEI Renjun, MATAMOROS E, LIU Manhong, et al. Training a molecular automaton to play a game[J]. Nature Nanotechnology, 2010, 5(11): 773–777. doi: 10.1038/nnano.2010.194
|
|
SONG Tianqi, GARG S, MOKHTAR R, et al. Analog computation by DNA strand displacement circuits[J]. ACS Synthetic Biology, 2016, 5(8): 898–912. doi: 10.1021/acssynbio.6b00144
|
|
QIAN Lulu and WINFREE E. Scaling up digital circuit computation with DNA strand displacement cascades[J]. Science, 2011, 332(6034): 1196–1201. doi: 10.1126/science.1200520
|
|
SONG Tianqi, ESHRA A, SHAH S, et al. Fast and compact DNA logic circuits based on single-stranded gates using strand-displacing polymerase[J]. Nature Nanotechnology, 2019, 14(11): 1075–1081. doi: 10.1038/s41565-019-0544-5
|
|
QIAN Lulu, WINFREE E, and BRUCK J. Neural network computation with DNA strand displacement cascades[J]. Nature, 2011, 475(7356): 368–372. doi: 10.1038/nature10262
|
|
BUI H, SHAH S, MOKHTAR R, et al. Localized DNA hybridization chain reactions on DNA origami[J]. ACS Nano, 2018, 12(2): 1146–1155. doi: 10.1021/acsnano.7b06699
|
|
LIU Huajie, WANG Jianbang, SONG Shiping, et al. A DNA-based system for selecting and displaying the combined result of two input variables[J]. Nature Communications, 2015, 6: 10089. doi: 10.1038/ncomms10089
|
|
CHATTERJEE G, DALCHAU N, MUSCAT R A, et al. A spatially localized architecture for fast and modular DNA computing[J]. Nature Nanotechnology, 2017, 12(9): 920–927. doi: 10.1038/nnano.2017.127
|
|
BOEMO M A, LUCAS A E, TURBERFIELD A J, et al. The formal language and design principles of autonomous DNA walker circuits[J]. ACS Synthetic Biology, 2016, 5(8): 878–884. doi: 10.1021/acssynbio.5b00275
|
|
SHERMAN W B and SEEMAN N C. A precisely controlled DNA biped walking device[J]. Nano Letters, 2004, 4(7): 1203–1207. doi: 10.1021/nl049527q
|
|
YIN Peng, YAN Hao, DANIELL X G, et al. A unidirectional DNA walker that moves autonomously along a track[J]. Angewandte Chemie: International Edition, 2004, 43(37): 4906–4911. doi: 10.1002/anie.200460522
|
|
BATH J, GREEN S J, and TURBERFIELD A J. A free-running DNA motor powered by a nicking enzyme[J]. Angewandte Chemie: International Edition, 2005, 44(28): 4358–4361. doi: 10.1002/anie.200501262
|
|
LUND K, MANZO A J, DABBY N, et al. Molecular robots guided by prescriptive landscapes[J]. Nature, 2010, 465(7295): 206–210. doi: 10.1038/nature09012
|
|
WICKHAM S F J, BATH J, KATSUDA Y, et al. A DNA-based molecular motor that can navigate a network of tracks[J]. Nature Nanotechnology, 2012, 7(3): 169–173. doi: 10.1038/nnano.2011.253
|
|
GU Hongzhou, CHAO Jie, XIAO Shoujun, et al. A proximity-based programmable DNA nanoscale assembly line[J]. Nature, 2010, 465(7295): 202–205. doi: 10.1038/nature09026
|
|
JUNG C, ALLEN P B, and ELLINGTON A D. A stochastic DNA walker that traverses a microparticle surface[J]. Nature Nanotechnology, 2016, 11(2): 157–163. doi: 10.1038/nnano.2015.246
|
|
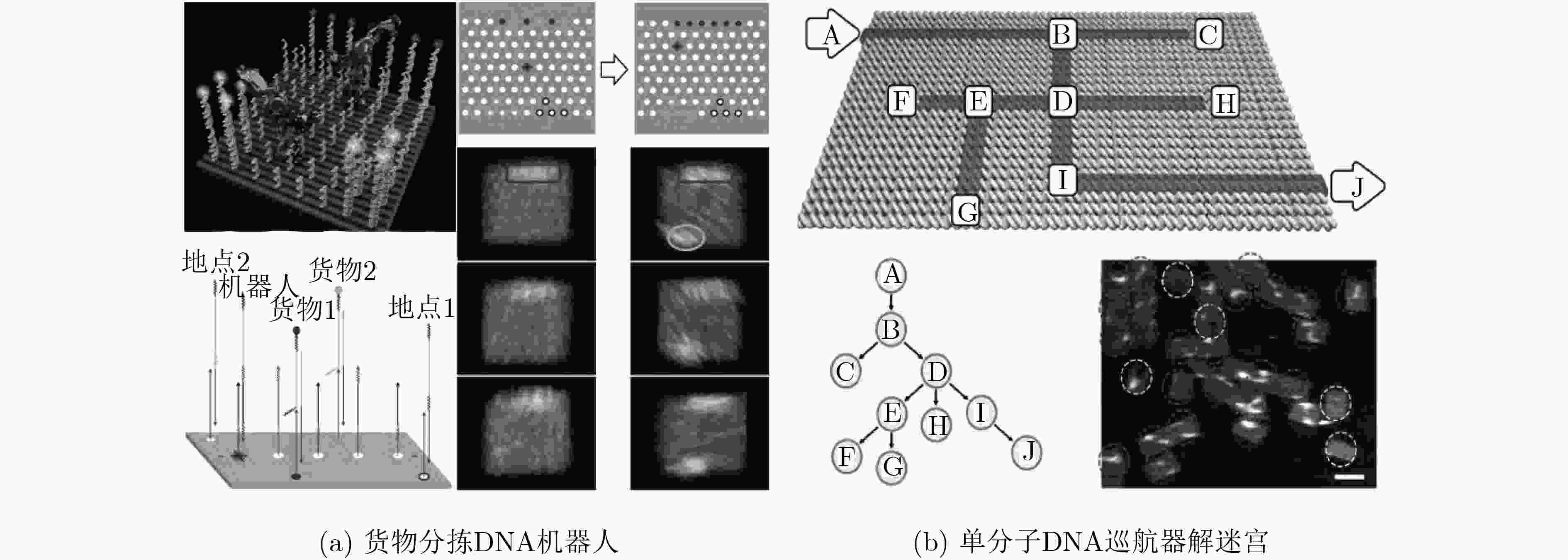
THUBAGERE A J, LI Wei, JOHNSON R F, et al. A cargo-sorting DNA robot[J]. Science, 2017, 357(6356): eaan6558. doi: 10.1126/science.aan6558
|
|
CHAO Jie, WANG Jianbang, WANG Fei, et al. Solving mazes with single-molecule DNA navigators[J]. Nature Materials, 2019, 18(3): 273–279. doi: 10.1038/s41563-018-0205-3
|
|
DIRKS R M and PIERCE N A. Triggered amplification by hybridization chain reaction[J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(43): 15275–15278. doi: 10.1073/pnas.0407024101
|
|
SCHULMAN R, YURKE B, and WINFREE E. Robust self-replication of combinatorial information via crystal growth and scission[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(17): 6405–6410. doi: 10.1073/pnas.1117813109
|
|
WOODS D, DOTY D, MYHRVOLD C, et al. Diverse and robust molecular algorithms using reprogrammable DNA self-assembly[J]. Nature, 2019, 572(7771): E21. doi: 10.1038/s41586-019-1378-x
|
|
JINEK M, CHYLINSKI K, FONFARA I, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity[J]. Science, 2012, 337(6096): 816–821. doi: 10.1126/science.1225829
|
|
CURRIN A, KOROVIN K, ABABI M, et al. Computing exponentially faster: Implementing a non-deterministic universal turing machine using DNA[J]. Journal of the Royal Society Interface, 2017, 14(128): 20160990. doi: 10.1098/rsif.2016.0990
|





 下载:
下载:


 下载:
下载:
